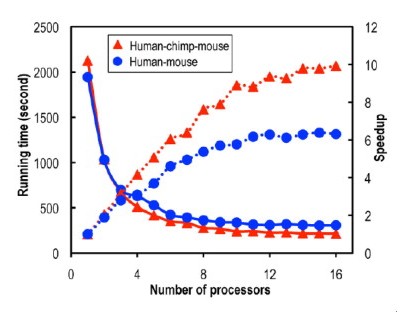
A parallel tool for constructing multiple protein-coding DNA alignments called ParaAT (Parallel Alignment & back-Translation) was recently created by Professor ZHANG Zhang and his team at Beijing Institute of Genomics (BIG), Chinese Academy of Sciences (CAS). Protein-coding DNA alignments have been studying by bioinformaticians to unveil the mystery of homologous sequences within and among species. It is pointed by Prof. ZHANG that “back-translated nucleotide alignments guided by amino acid alignments are more reliable and accurate than direct nucleotide alignments”. The traditional way for protein-coding nucleotide alignments is to construct an alignment based on protein sequences and then back-translate into the corresponding coding DNA alignment, which only accepts one homologous group as input and has limitations of generating multiple protein-coding DNA alignments for a large number of homologous groups. To catch up with the fast expending high-throughput era, a parallel tool ParaAT was created and developed. It parallelly aligns protein sequences and back-translates the aligned sequences into the corresponding DNA alignments, and thus is capable of constructing protein-coding DNA alignments for a large number of homologs. ParaAT is provided as open-source software and can be executed on Linux/Unix, Macintosh, and Windows platforms. The ParaAT package, including source codes, example data and documentation, is freely available for academic use at http://cbb.big.ac.cn/software. Paper link: http://www.sciencedirect.com/science/article/pii/S0006291X12003518?v=s5 Speedup (dotted lines) and running time (solid lines) for constructing protein-coding DNA alignments using 1–16 CPUs. (Image by ZHANG zhang) Contact: Prof. ZHANG Zhang Email: zhangzhang@big.ac.cn