CancerSCEM: A Database of Single-cell Expression Map Across Various Human Cancers
Single-cell transcriptome sequencing (scRNA-seq), which has significant advantages in studying cell heterogeneity at the single cell resolution, has become an indispensable method for studying the tumor microenvironment, cancer pathogenesis, metastasis and invasion, and the treatment and diagnosis of various cancers.
Until November 2021, more than 1,300 cancer-related single-cell transcriptome studies were included in PubMed, which has greatly improved our understanding of cancer initiation and development, and also promoted the clinical diagnosis and treatment of diverse cancers. The exponential explosion of massive cancer scRNA-seq data during the past decade are calling for a burning demand to be integrated and processed for essential investigations in tumor microenvironment.
In response to this demand, the research group of the National Genomics Data Center, Beijing Institute of Genomics of Chinese Academy of Sciences/ China National Center for Bioinformation (CNCB) developed CancerSCEM,a database of Cancer Single-cell Expression Map,this work was published online in Nucleic Acid Research with the title “CancerSCEM: a database of single-cell expression map across various human cancers”.
CancerSCEM consists of 208 cancer scRNA-seq datasets across 20 human cancer types, such as lung adenocarcinoma (LUAD), colorectal cancer (CRC)and glioblastoma (GBM). A standardized analysis workflow was built and accurate cell-type annotation was obtained for each individual sample. A series of additional analyses were performed which provide users with more abundant tumor microenvironment related information, such as differential gene expression analysis, expression profiling of receptor-ligand gene pairs, cell-cell interaction network construction et al.
Furthermore, survival analysis based on TCGA expression data and clinical information was also carried out.
CancerSCEM provides mainly 4 function modules: browsing, searching, online analyzing and downloading. Users can adopt quick search, keyword cloud or advanced search for querying cancer scRNA-seq datasets or samples of interest.36 widely-used immune checkpoint molecules (such as PDCD1, TIGIT and LAG3) were curated and provided as a special list in the database, researchers and clinicians thereby can directly access to their expression profiles across different cancer types, and quickly narrow down the optimal targets for clinical immunotherapies.
Moreover, CancerSCEM is also equipped with an interactive comprehensive online analysis platform, which includes 2 modules and 7 functions. Users are able to perform multi-level real-time analyses: Comparison of gene expression in different cell types, Cell composition comparison between samples, Cell interaction network construction and Survival analysis, etc. This analysis platform will undoubtedly provide users with friendly services for personalized cancer scRNA-seq data mining.
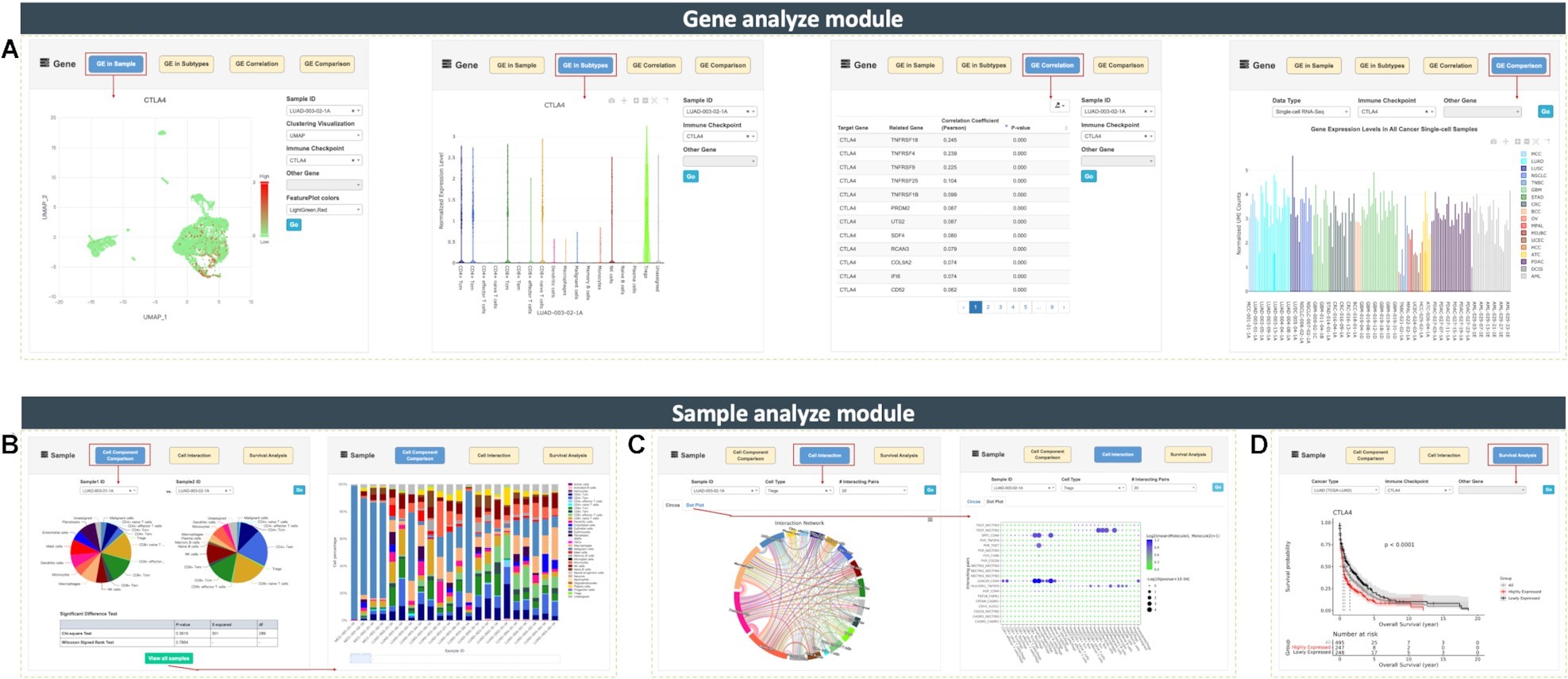
Screen captures of seven online analyzing functions equipped in CancerSCEM (Image by XIAO Jingfa's group)
Contact:
Dr. XIAO Jingfa
E-mail: xiaojingfa@big.ac.cn
National Genomics Data Center (NGDC)