scMethBank: A Database for Single-cell Whole Genome DNA Methylation Maps
As an important part of epigenomics, DNA methylation provides an important perspective for studying biological processes including genomic imprinting, early embryo development, and cancer progression. With the continuous development of single-cell sequencing technology, advances in sequencing methods have enabled the development of strategies to analyze DNA methylation at single-cell resolution, which greatly promotes the exploration of cell epigenetic heterogeneity.
However, the continuous accumulation of the vast amount of experiments and datasets poses a great challenge for the integration and reuse of single-cell DNA methylation data. Despite some efforts, there is still a drastically short of a systematic single-cell DNA methylation database up to now.
To address these challenges, the research group of the National Genomics Data Center, Beijing Institute of Genomics of Chinese Academy of Sciences/ China National Center for Bioinformation (CNCB) developed scMethBank. And this work has been published online in Nucleic Acid Research with the title “scMethBank: A database for single-cell whole genome DNA methylation maps”.
By collecting data produced by several kinds of main bisulfite-based strategies and characterizing methylation status at single-base resolution with a standardized workflow, scMethBank builds the whole genome methylation maps at the single-cell level for human and mouse. It provides the genome-wide single-cell DNA methylation profiles and value-added curation metadata across multiple biological conditions including cell types, developmental stages, disease states, and treatment methods. Moreover, random access to methylation data of any genome interval has been achieved by optimizing the underlying database storage and inquiry. A user-friendly interface is equipped to support interactive exploration and visualization of single-cell methylation profiles with single-base accuracy.
Other functions in scMethBank include data searching and downloading, interactive genome browser with custom tracks, and online analysis and graphical tools specifically designed for the single-cell methylation data. We expect this database can provide valuable single-cell methylation data resources to accelerate scientific discoveries and lead to a new paradigm into epigenetic heterogeneity.
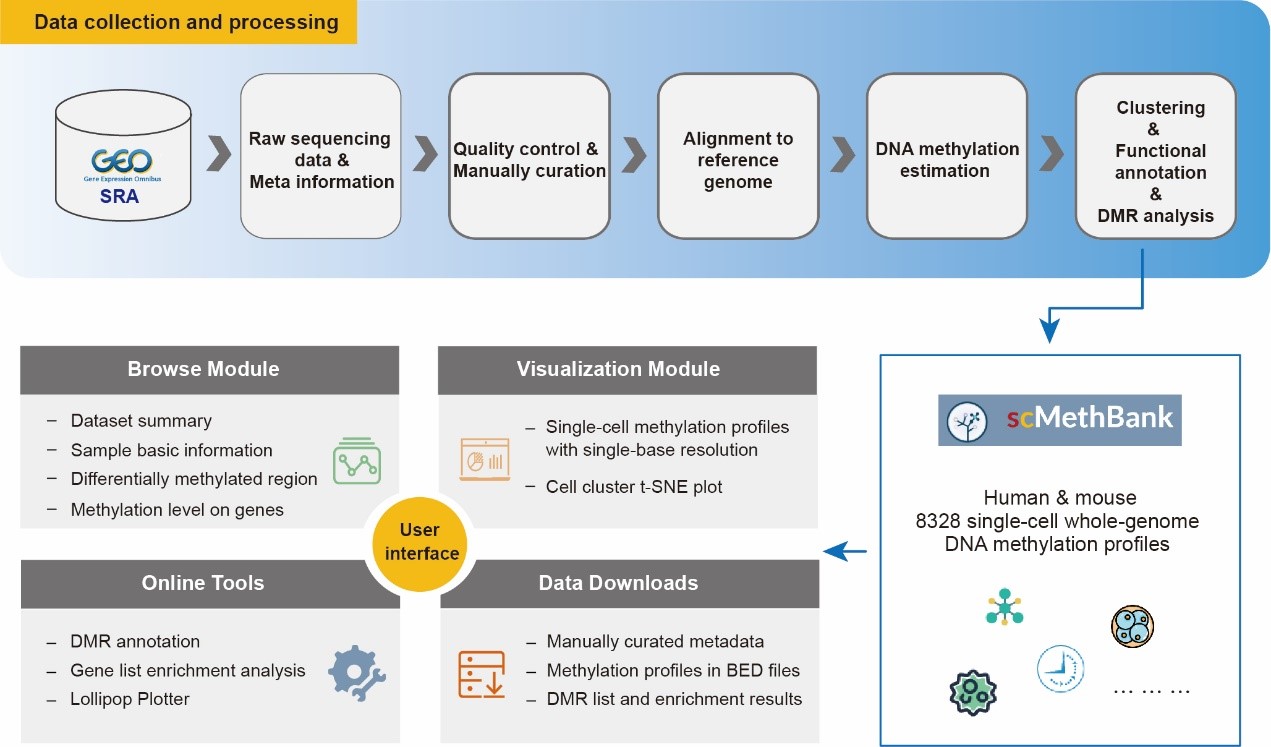
Database contents and features of scMethBank (Image by NGDC)
Contact:
Dr. LI Rujiao
Email: lirj@big.ac.cn
Dr. BAO Yiming
Email: baoym@big.ac.cn