EWAS Open Platform: Integrated Data, Knowledge and Toolkit for Epigenome-wide Association Study
With the explosive growth of epigenome-wide association studies (EWAS), huge amounts of data and knowledge related to EWAS have been accumulated. Since these data hold great potential for clinical translations, a standardized platform for data archive, retrieval and exploration is indispensable. In 2019 and 2020, the research group of the National Genomics Data Center (NGDC), Beijing Institute of Genomics of Chinese Academy of Sciences/ China National Center for Bioinformation (CNCB) has developed EWAS Atlas (Nucleic Acid Research, 2019) and EWAS Data Hub (Nucleic Acid Research, 2020).
Since the first release of EWAS Atlas, there are over 33,000 visitors with a total of 147,000 accesses, along with >100 emails and phone calls from worldwide users. In order to provide a comprehensive system for EWAS data storage and download, knowledge collection and browsing, and downstream analysis and visualization, the research group of NGDC developed EWAS Open Platform, a database of integrated data, knowledge and toolkit for epigenome-wide association study, which was published online in Nucleic Acid Research.
EWAS Open Platform is an open platform for epigenome-wide association studies that incorporates three components: EWAS Data Hub for data collection and standardized normalization, EWAS Atlas for knowledge extraction and curation, and EWAS Toolkit for downstream analysis and visualization. Each component is a stand-alone database or web server.
EWAS Data Hub integrates a comprehensive collection of DNA methylation array data from 115,852 samples and employs an effective normalization method (GMQN) to remove batch effects among different datasets. Accordingly, taking advantages of both massive high-quality DNA methylation data and standardized metadata, EWAS Data Hub provides reference DNA methylation profiles under different contexts.
EWAS Atlas integrates a large number of 617,018 high-quality EWAS associations, involving 193 tissues/cell lines and covering 619 traits and 3385 cohorts, which are completely based on manual curation from 1440 studies reported in 910 publications.
As an indispensable component of EWAS Open Platform, EWAS Toolkit is a new powerful one-stop analysis service for EWAS downstream analysis. Currently, EWAS Toolkit integrates knowledge and data, organically combines EWAS Atlas and EWAS Data Hub, and provides users with a wide range of analysis and visualization including enrichment, annotation and network visualization.
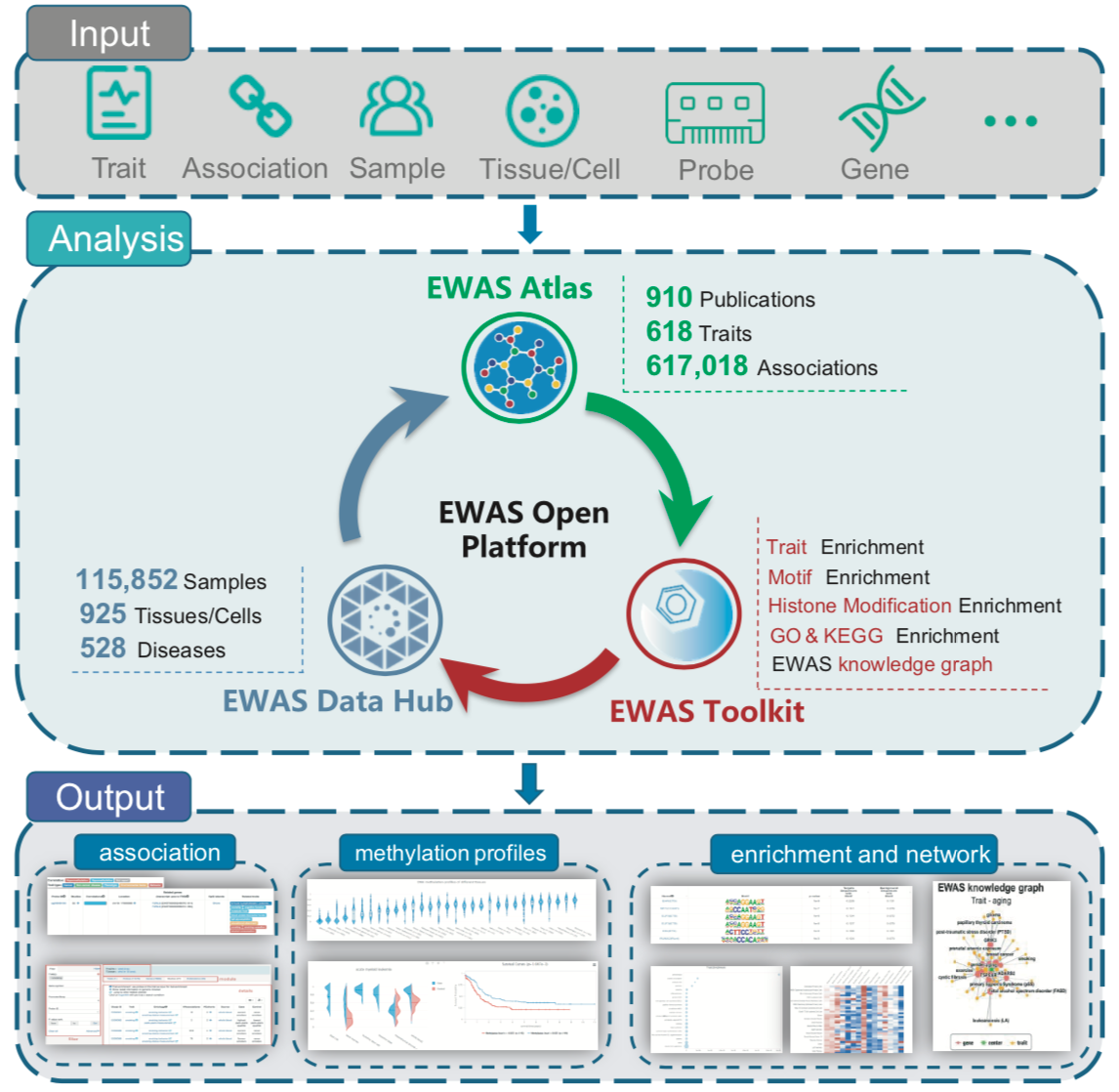
Schematic overview of EWAS Open Platform data processing workflow (Image by NGDC)
Contact:
Dr. LI Rujiao
Email: lirj@big.ac.cn
Dr. ZHANG Zhang
Email: zhangzhang@big.ac.cn
Dr. BAO Yiming
Email: baoym@big.ac.cn