GWAS Atlas: An Updated Knowledgebase Integrating More Curated Associations in Plants and Animals
Genome-wide association study (GWAS) is a key technique for exploiting the genetic basis of complex traits and diseases. The genotype-phenotype association (G2P) identified by GWAS is an important resource to reveal the molecular mechanism of complex traits.
Recently, the National Genomics Data Center, Beijing Institute of Genomics of CAS (China National Center for Bioinformation) present an updated resource entitled “GWAS Atlas: a curated resource of genome-wide variant-trait associations in plants and animals” for the
Nucleic Acids Research 2022 Database Issue.
The GWAS Atlas team has set up a standardized curation process to curate publicly available G2Ps. Specifically, the current release of
GWAS Atlas incorporates a curated collection of 278,109 G2Ps for 15 species (10 plants and 5 animals) from 830 publications. To prioritize putative causal variants for further study, we perform lead SNP analysis to select significantly independent SNPs, resulting in 6,084 lead SNPs. In addition, we curate a comprehensive collection of 486 experimentally identified causal variants associated with 157 traits. Together, the integrated knowledge is of great usefulness for better understanding the genetic architecture of complex traits and for aiding precise designing and breeding.
To improve services regarding data submission and analysis, GWAS Atlas offers online data submission services and allows GWAS results submission for any species. Each submission will be assigned a unique identifier and be released according to the date set by the submitter. Besides, GWAS Atlas also provides four commonly used tools, including LeadSNPFinder, GeneFinder, MHPlotter, and QQPlotter, which can facilitate users to perform data mining and visualize GWAS results. In conclusion, GWAS Atlas provides an important data management and analysis platform for agronomic trait study and molecular breeding.
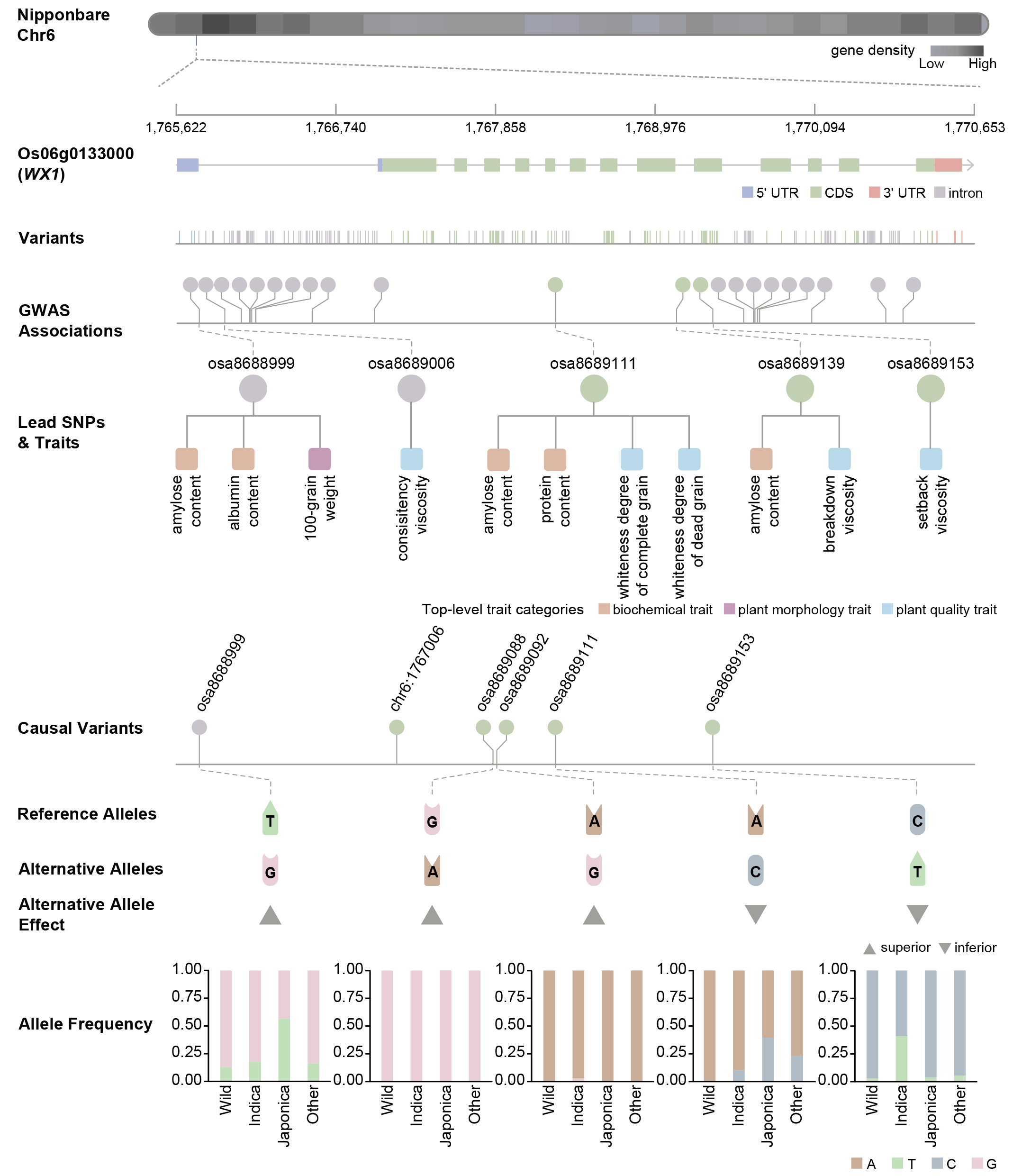
Variants in WX1 gene. Variants in WX1 are organized as polymorphic variants, GWAS associated variants, lead SNPs identified by computational methods, and causal variants curated from publication (Image by NGDC).
Contact:
Dr. ZHANG Zhang
Email: zhangzhang@big.ac.cn