RCoV19: One-stop Hub for SARS-CoV-2 Genome Data Integration, Variant Monitoring, and Risk Pre-warning
The coronavirus disease 2019 (COVID-19) is the most extensive and consequential epidemic in nearly a century. The number of genomic sequences of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is far exceed the sum of sequences of other known viruses. The vast number of genome sequences presents unprecedented challenges for rapid integration, analysis, and data mining. The COVID-19 pandemic is still spreading globally, and the genome of SARS-CoV-2 is constantly mutating and evolving. Therefore, developing and establishing a platform for automated data integration, real-time monitoring, and high-risk variant pre-warning of large-scale SARS-CoV-2 genomic data has important application value and scientific significance.
The Beijing Institute of Genomics of the Chinese Academy of Sciences (China National Center for Bioinformation) launched the first open-access resource of SARS-CoV-2, RCoV19, on January 22, 2020. RCoV19 is constantly updating and incorporating global genomic sequences and metadata of SARS-CoV-2. It supports the storage, sharing, and convergence of SARS-CoV-2 genome sequences, as well as providing mutation annotation information and evolutionary lineage data. This platform has become the largest and most comprehensive public resource for SARS-CoV-2 research worldwide.
To better serve scientific research on SARS-CoV-2 and the construction of genomic big data-based monitoring and pre-warning systems, research team comprehensively upgraded and improved RCoV19, which has significant improvements and advancements compared with the previous version. This work was published online in the journal Genomics, Proteomics & Bioinformatics.
Firstly, RCoV19 has developed a fully automated data intelligent curation model and data sharing platform for automating the collection, de-redundancy, cross-referencing, and quality assessment of SARS-CoV-2 genome data. This model continuously provides integration of sequences, metadata, global distribution, and statistics of SARS-CoV-2 in real time, along with efficient personalized advanced search services.
Secondly, with the integration of massive data from RCoV19, they have developed a rapid variation analysis method for genome sequences, an algorithm for constructing haplotype network evolution, and a high-risk variant pre-warning model. These tools have enabled researchers to create a real-time monitoring platform for the transmission and evolution of SARS-CoV-2, a visual pre-warning system for high-risk variants, and an interactive quick comparison module for mutation spectra. Using these tools, researchers can dynamically visualize the genomic sequences, variations, and evolutionary lineages of SARS-CoV-2.
Additionally, RCoV19 can provide early warnings for high-risk variants and analyze the characteristics of important sequences or lineages. It offers critical technical and data support for public health security responses driven by genomic big data.
Lastly, RCoV19 curated some effect knowledge of SARS-CoV-2 genome mutation, which includes the impact on infectivity/transmissibility, antibody resistance, drug resistance, and T-cell epitopes. This information assists researchers and policymakers in comprehending the mutation characteristics of SARS-CoV-2 more effectively, and provides a crucial reference for scientific research and decision-making related to prevention and control measures.
In summary, RCoV19 is a comprehensive platform that can automatically integrate SARS-CoV-2 genomic data, conduct mutation monitoring, provide high-risk variant pre-warnings and mutation effect knowledge. It aims to promote scientific research on SARS-CoV-2 and provide strong support for the establishment of the global public health security system.
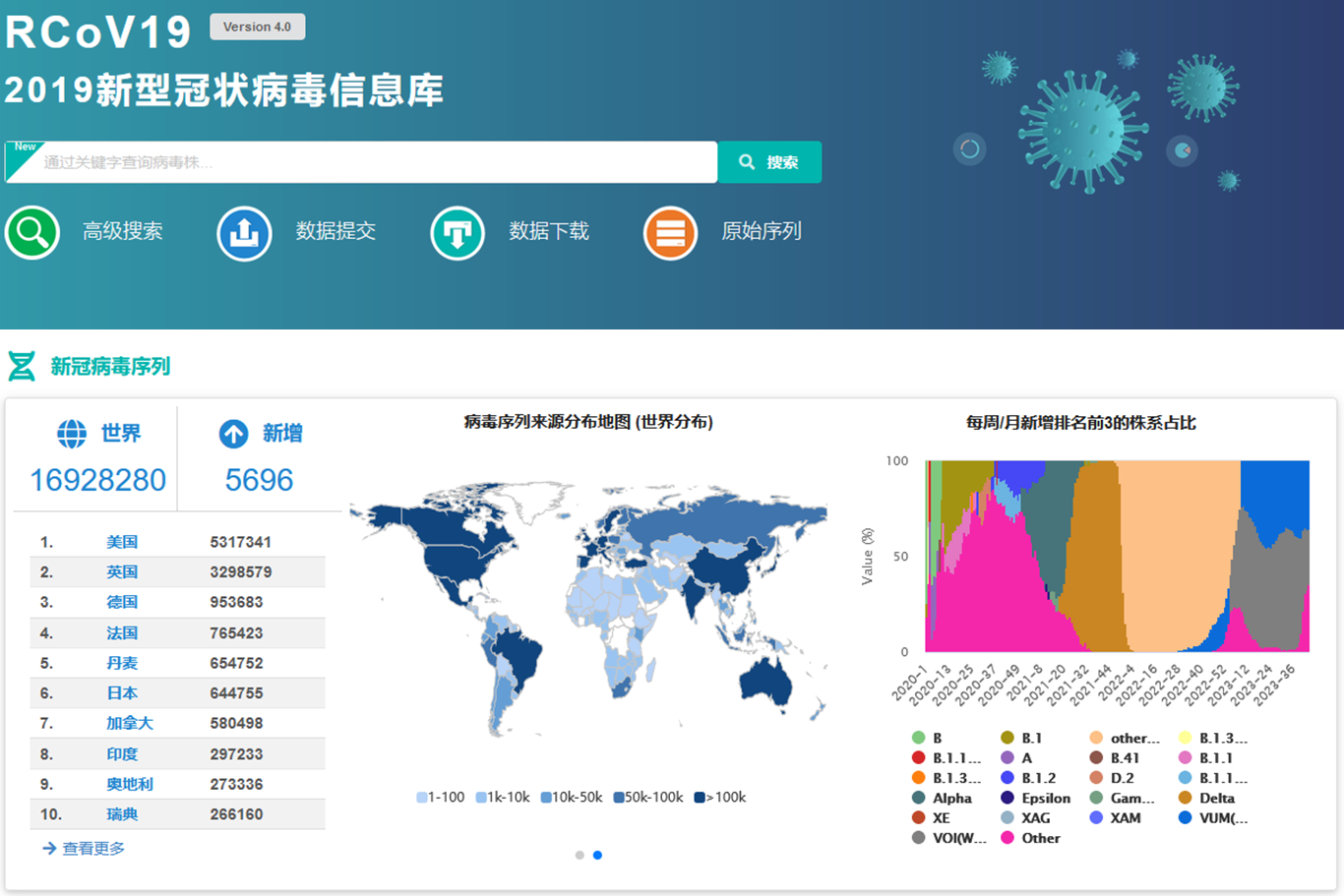
Homepage of RCoV19
Contact:
Dr. BAO Yiming
Email: baoym@big.ac.cn
Dr. SONG Shuhui
Email: songshh@big.ac.cn