N6-methyladenosine Regulates RNA Abundance of SARS-CoV-2
The worldwide pandemic of COVID-19 is caused by a novel β-coronavirus SARS-CoV-2, an enveloped RNA virus with around 30 kb of a positive-sense, single-stranded RNA genome. In the host cells, the SARS-CoV-2 RNA firstly synthesizes the negative-sense RNA using its RNA-dependent RNA polymerase (RdRP), and then uses the negative-sense RNA as a template to produce positive-sense genomic and subgenomic RNAs.
RNA modifications have been identified to be present in both genomic and subgenomic RNAs of SARS-CoV-2. As the most abundant RNA methylation in mRNA, N6-methyladenosine (m6A) involves in various RNA processing and plays important roles in post-transcriptional regulations. However, the precise landscape of m6A in SARS-CoV-2 and its potential significance in regulating viral RNA metabolism and lifecycle remain unknown.
A recent study led by Dr. YANG Yungui’s lab from Beijing Institute of Genomics of the Chinese Academy of Sciences (China National Center for Bioinformation) and Dr. WANG Jianwei’s lab from Institute of Pathogen Biology of the Chinese Academy of Medical Sciences uncovered the key role of m6A in regulating abundance of SARS-CoV-2 RNA. It has been published in Cell Discovery.
The researchers first used the specific m6A antibody and performed MeRIP-seq to obtain the high-resolution m6A landscape of SARS-CoV-2 RNA from infected Vero cells. They found that the depletion of m6A methyltransferase METTL3 could increase the abundance of viral RNA in host cells, suggesting that m6A might play an important regulatory role in metabolism of viral RNA.
Then, the researchers used junction reads to quantify the expression levels of each subgenomic RNA. They found that the expression of not some specific subgenomic RNAs but all of them increased upon depletion of METTL3, indicating that there was a conserved m6A-dependent mechanism for regulating all types of SARS-CoV-2 RNAs.
Besides, the researchers quantified methylation level for each m6A site between normal and METTL3 knockdown Vero cells. They found that the METTL3-targeted m6As were enriched around 3’ termini shared by all types of viral transcripts, suggesting that these m6As might involve in regulation of viral RNA degradation. More intriguingly, viral transcripts could evade this regulation by formation of shorter 3’UTRs without m6A sites through alternative splicing.
This study reveals the m6A methylome of SARS-CoV-2 and points to the significance of m6A in regulating SARS-CoV-2 RNA metabolism in host cells. It also provides theoretical and technical supports for further investigation on the underlying mechanism and biological functions.
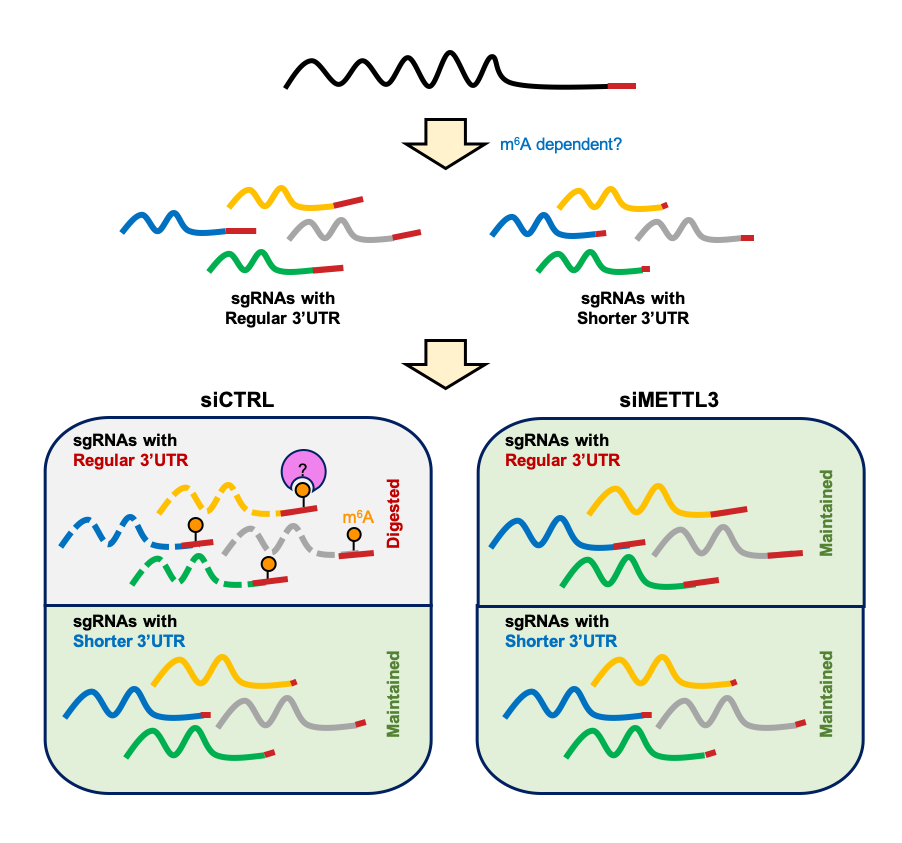
Schematic model of the m6A regulating RNA abundance of SARS-CoV-2 (Image by YANG's group)
Contact:
Dr. YANG Yungui
Email: ygyang@big.ac.cn